Demystifying DNA 2: Y-DNA tests
Originally posted 1st July 2018, updated 4th October 2024.
In my last blog we looked at the science behind DNA testing and the different types of test available. In these next few posts in the series we will look at each type of DNA testing in more detail.
Uses of Y-DNA
Y-DNA is passed from father to son, unchanged for many generations, and this makes it a powerful tool for assessing your paternal line: a match on a Y-DNA test can only lead you up one part of your family tree.
Y-DNA is often used by those running surname studies as, in principle, the descent of the male line is the same as the descent of the surname. Y-DNA can therefore be used to assess the likelihood of all bearers of a particular surname arising from the same single individual, no matter how far back in time this individual lived. There are single surname DNA projects through the commercial testing sites and a number of One-Name Studies (ONS) also operate DNA projects.
However, surnames do not always reflect bloodlines. The most common issue is the bearer of a surname being found to be illegitimate: Mr Postlethwaite was actually a plain old Mr Brown. In DNA circles this is referred to as a “non-paternity event” or "not the parent expected" (NPE), or "misattributed parent event" (MPE). There are a number of other reasons a surname may be assumed: unofficial adoption, taking on a stepfather’s surname and so on.
Y-DNA can also be used to identify an unknown father, though this use has been largely replaced with autosomal DNA tests now for recent generations. The caveat for Y-DNA is that the results may point to a particular male line but not necessarily an individual within that line. Just using Y-DNA testing could point to Mr A being the father, but equally his brother, his paternal cousin or his uncle. As with all types of DNA test, the DNA data is just one source of information that can be used to aid genealogical research. It needs to be used in the context of known information and documentary research, in this case, who was in the right place at the right time?
Who can take a test?
Y-DNA tests can only be taken by males. However, the power of the Y-DNA chromosome is that it is unchanged over many generations. If you have no brothers you can ask a cousin to test, or an uncle, or even a second or third cousin, So long as they are from an unbroken line of males from your common ancestor. The Y-DNA carrying descendants of an individual are illustrated below:
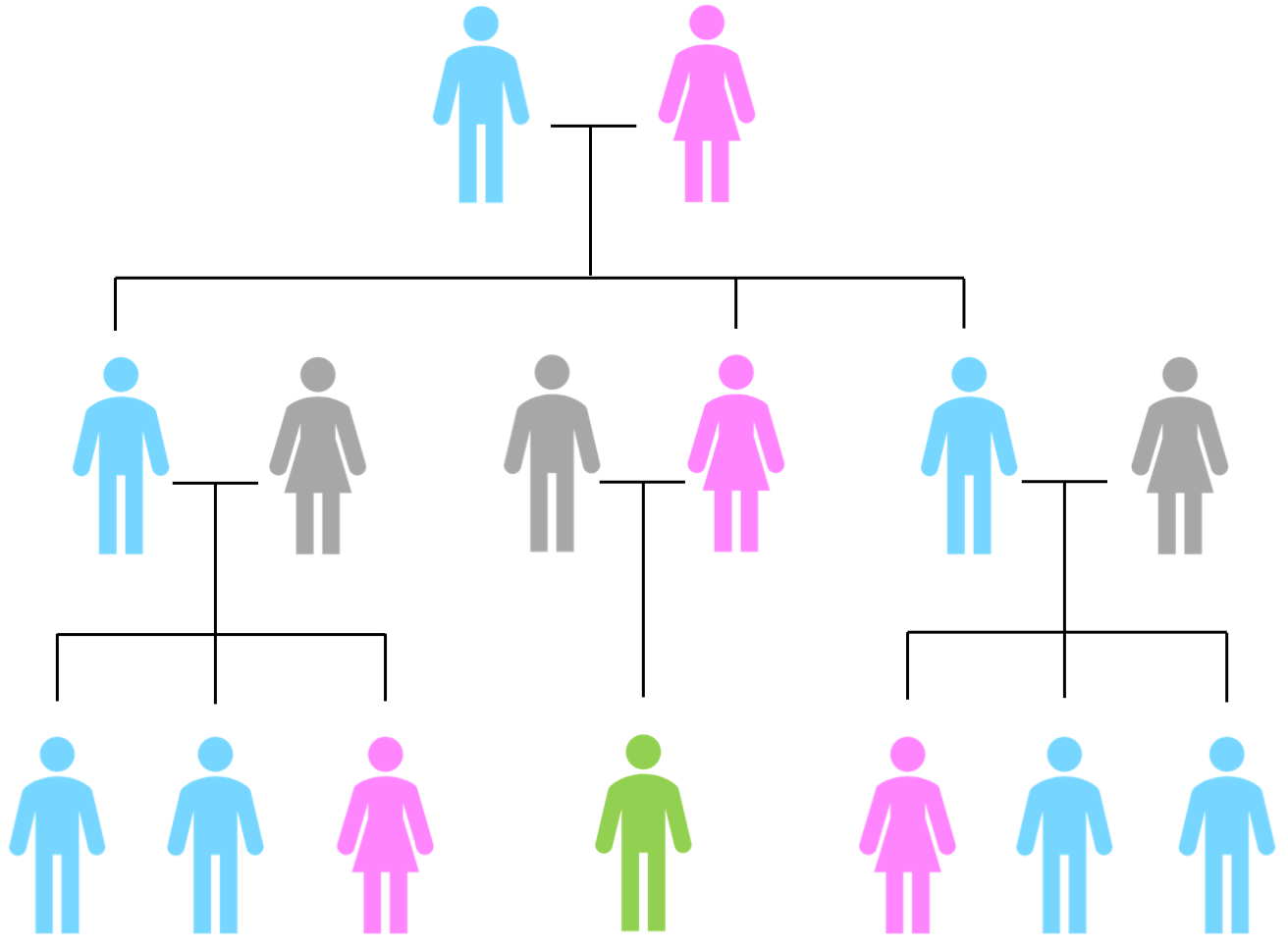
In the diagram, the descendants of a couple are shown. They have three children: two boys and a girl. The spouses of these three children are shown in grey to indicate they are not blood relatives of the couple at the top. The male descendants of the uppermost male, who descend through the male to male line and therefore carry Y-DNA, are coloured in blue. In the middle is a male filled in in green to show that, whilst he descends from the uppermost male, he descends via a daughter and therefore does not carry the same Y-DNA as those shown in blue i.e. the chain of descent for this green male is not male to male to male.
Let's assume that this green male is the genealogist of the family and he wants to know more about his maternal grandfather's patrilineal (male to male) line. Unfortunately, his grandfather died some years ago. Looking at the diagram you can see that he has options to ask both uncles and first cousins to take a test to represent his grandfather's Y-DNA. What if there are no surviving cousins or uncles to test? The joy of Y-DNA is that it remains unchanged for many generations, so you could go back another generation to our genealogist's great grandfather, and look at other any male to male descendants in the form of second cousins, and so on.
Generally speaking, if you have a choice, test the older generation. This approach does assume that all those in blue are the blood line of the uppermost male. A non-paternity event is a possibility and you should always test more than one candidate from more than one line of descent if you can.
The Data
There are two different methods of analysing and comparing Y-DNA:
- STR or Short Tandem Repeat testing
- SNP or Single Nucleotide Polymorph testing
STR Testing
Short Tandem Repeats (STRs) are positions along the DNA molecule where the same sequence of nucleotides or bases is characteristically repeated a number of times. For example, if at a particular position of the DNA molecule the bases were AGTCAGTCAGTCAGTCAGTC, the sequence AGTC is repeated five times.
Each STR marker is named, typically in the format DYS391 (where D = DNA, Y = Y chromosome and S = (unique) segment). The number of times the sequence is repeated at each marker is counted and the results of the Y-DNA test take the format shown below:

This set of results is called an individual’s haplotype. When purchasing a Y-DNA test you will see notations like Y-37, Y-67 and Y-111. These are the number of STR markers, or locations on the DNA molecule, tested. The example above tested at 12 markers only (there are two results at DYS385). Whilst early tests could only look at 12 markers, 37, 67 and 111 markers then became more common as the technology has developed, with some tests looking at even greater numbers of markers. Family Tree DNA now only sells the Y-37 and Y-111 tests (plus the Big Y, which we will come back to). You can still compare results with someone who tested with a different number of markers: a test at 37 markers looks at the same 12 markers as a 12 marker test, plus an additional 25, and so on.
The results of the STR tests are compared with those of others on the commercial websites to see if they are matches, i.e. have the same count of repeats at each marker tested. If all results match exactly the tester and match are said to have a genetic distance of zero. In the example above, if another set of results was compared and all were the same except the result for DYS391 was 9 instead of 10, this would be termed a genetic distance of one. If DYS391 was 9 and DYS426 was 13 this would be a genetic distance of 3, i.e. the difference of 1 at DYS391 plus the 2 on DYS426 (this is calculated in a slightly different way when you have more than one result at a marker, but this gives you the general idea).
The number of markers tested matters. If you have an exact match at 12 or 25 markers this does not necessarily mean you are closely related. A comparison of 67 markers is looking in more detail at the DNA and could reveal that there are actually large differences in the results. It's a bit like saying two men live in Yorkshire so are identical at 12 markers, but at 111 markers you are testing whether two men live at 49 Horton Grange Road, Bradford, Yorkshire.
Fortunately, you do not have to work any of this out yourself. At Family Tree DNA the
Time Predictor tool is used to predict when the likely
Most Recent Common Ancestor (MRCA) with a match lived, taking into account both the number of markers tested and the genetic distance value. The example below shows two men who have taken a Y-111 test and match with a genetic distance of 1. The Time Predictor report is access by clicking on the calendar icon highlighted in the image below. The Time Predictor report predicts the two individuals to share a common ancestor born in around 1850, with a range 1700-1900.
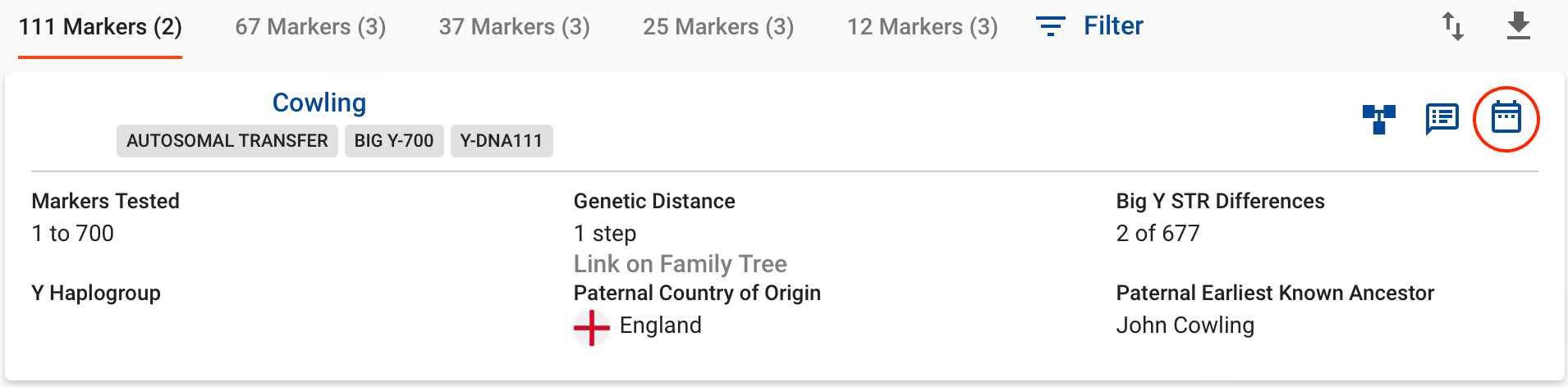
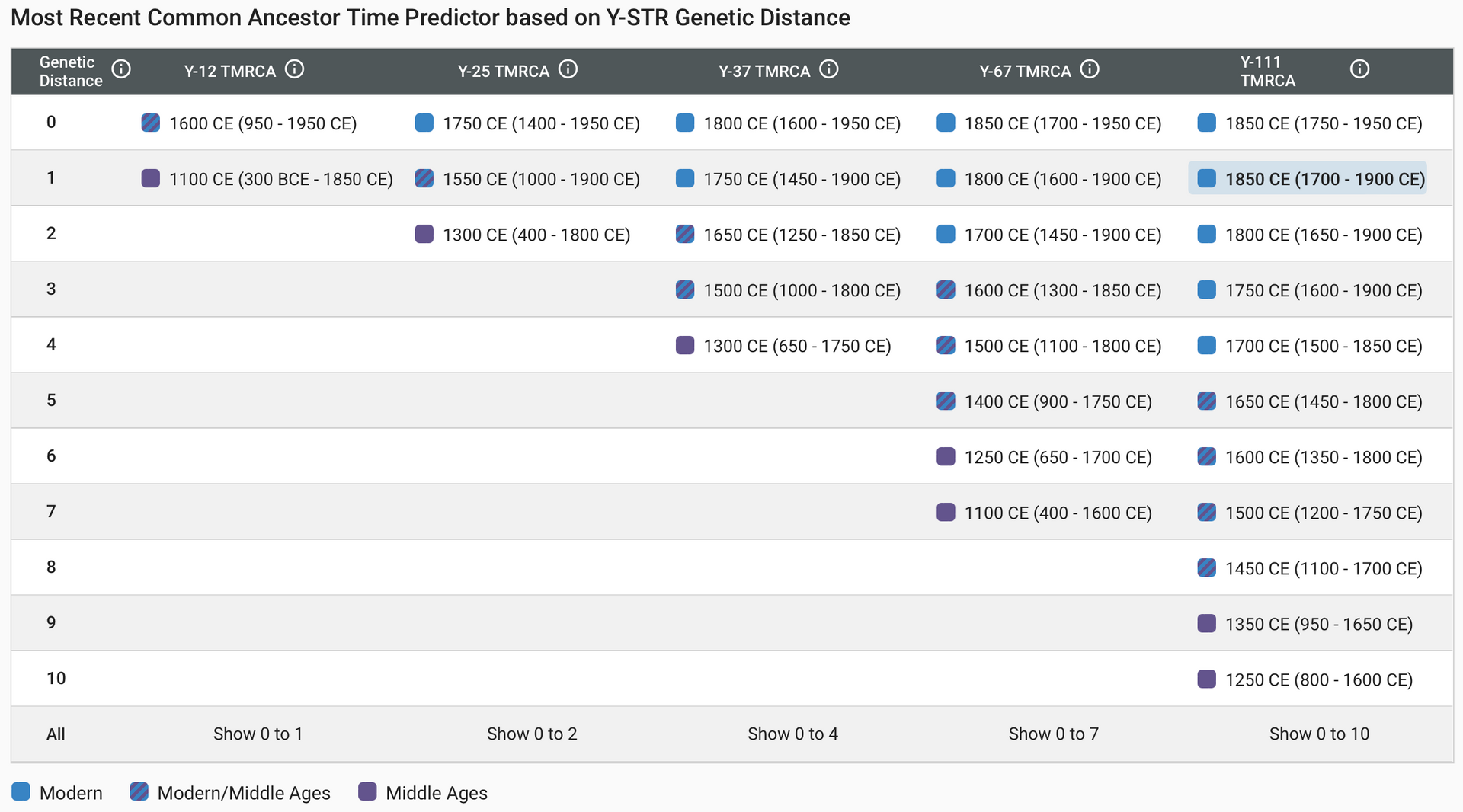
For many years STR testing was the backbone of matching with Y-DNA, and it certainly remains the most affordable type of testing. However, in recent years, the advances in technology and increasing numbers of those testing have meant that SNP testing, traditionally associated with looking at ancient origins and haplogroups, can also be useful in a genealogically relevant time frame.
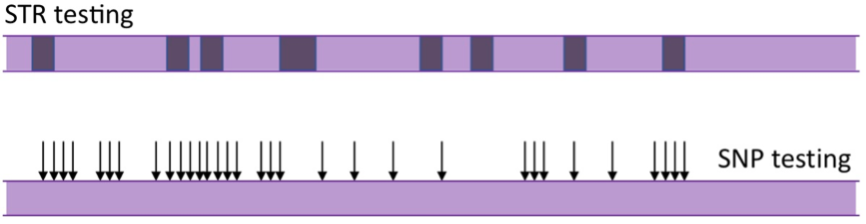
SNP testing
A Single Nucleotide Polymorph (SNP, pronounced “snip”) is a point along the DNA molecule known to differ from one individual to another – a point at which a mutation has occurred at some point in time. Rather than test areas of repeating nucleotide sequence like STR testing, this type of testing analyses which nucleotide is found at many individual locations or SNPs (pronounced “snips”).
Haplogroups and Mutations
An individual’s haplogroup is their location in the human Y-DNA haplogroup tree. Every male fits on this tree, some branches dating far further back in time than others. Traditionally, haplogroups were associated with looking at a male's ancient origins. A simple graphic is shown below but there are many branches, or subclades, within each haplogroup. The position of an individual on this "Y-DNA family tree" is dictated by whether or not they have mutations at particular SNP locations (i.e. have a different nucleotide, A, C, T, G, to other people). Going back to the address analogy: if two men live in Horton Grange Road they might be on the same branch, let's say haplogroup "I" on the image below. However, if one lives at number 49, for which a mutation has been identified, and the other at number 30, then only the 49 Horton Grange Road man moved down to the next branch or subclade, let's say "I1a", whilst the other man remains in haplogroup I.
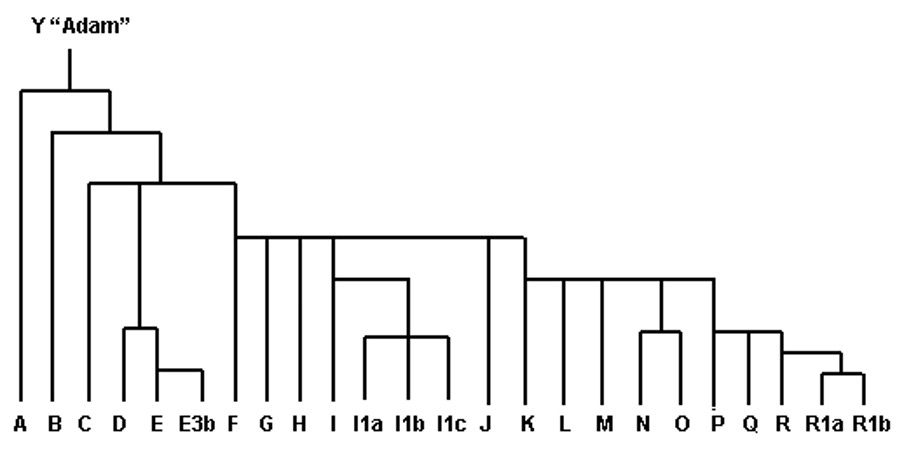
Y-DNA haplogroups used to be written in the form R1a1a1b2a2a1 but as more research was conducted more and more branches discovered this quickly became unwieldy. Now the name of the haplogroup is shortened to the letter from the major haplogroup branch, follow by the final SNP at which a mutation was detected. Care needs to be taken with comparing haplogroups, as one tester may have only an estimated haplogroup and another have had SNP testing conducted to a deeper level (i.e. they may appear different but could actually be from the same higher branch).
As technology has developed it has become possible to define more and more mutations and more and more branches, some of which were created in very recent generations. This means SNP testing can also be used to look at your DNA matches and how far back in time a common ancestor may have lived.
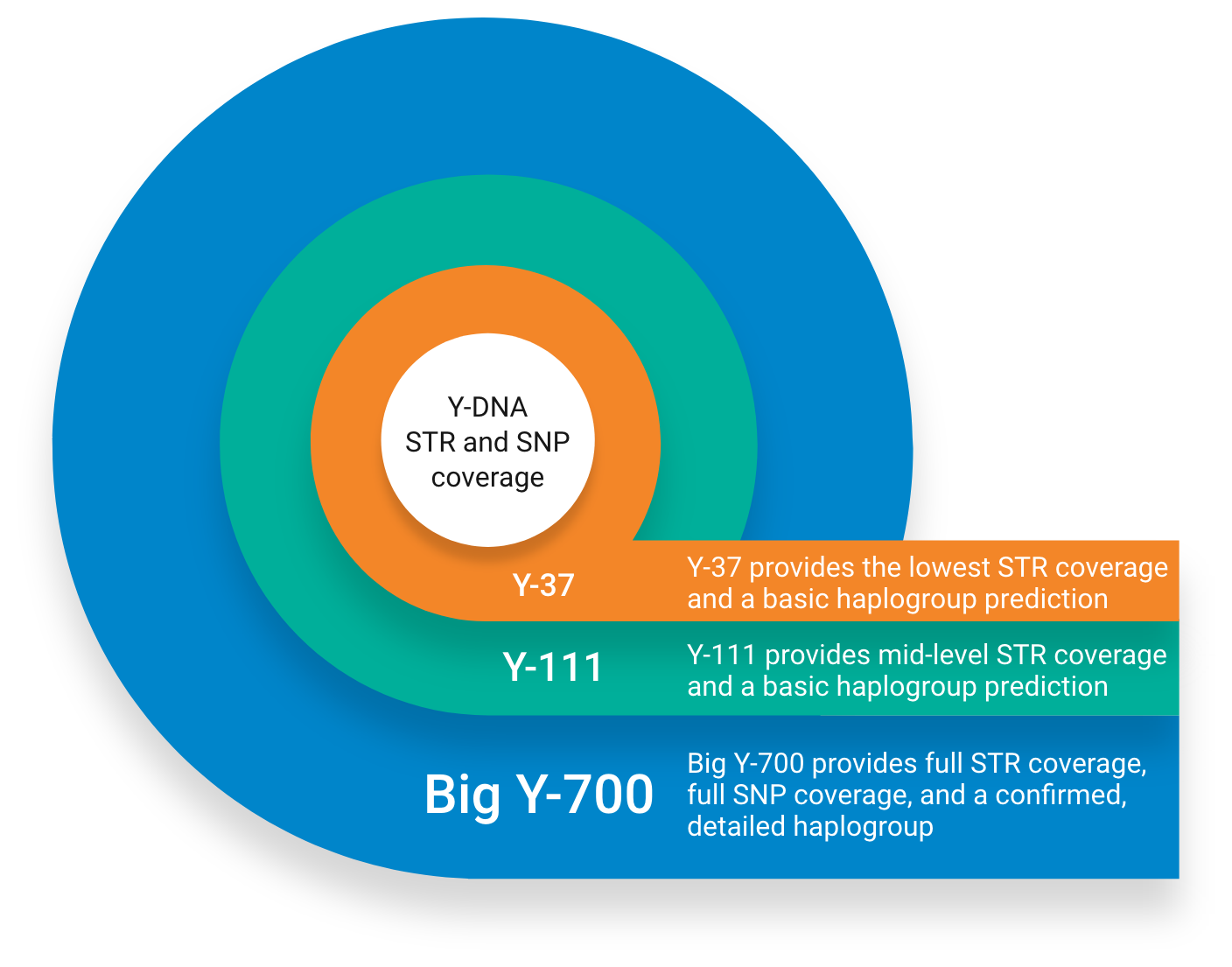
At Family Tree DNA this more detailed analysis of STRs and SNPs is called the Big-Y700 test, which tests over 700 STRs and hundreds and thousands of SNPs. Your results will include a more detailed estimate of your haplogroup than STR testing alone, and will include "private variants", mutations not yet classified as known SNPs. Ultimately, as more people test, your haplogroup will get further defined. It is the discovery and analysis of more recent mutations / variants that makes SNP testing useful for fine-tuning the time to MRCA for two individuals. However, at around $449 US dollars, this type of testing is for the more dedicated DNA enthusiast and those involved with surname and other Y-DNA projects. A more in-depth discussion of Big Y is beyond the scope of this introduction to Y-DNA. You can read more about the Big-Y test here. You can read my review of some of the latest Y-DNA tools you can use with Big Y tests here.
The Testing Companies
Only Family Tree DNA currently offers a separate Y-DNA test, where STR results can be compared with other users. Your own STR data can also be downloaded for further analysis elsewhere. Haplogroup and SNP testing is also available from Family Tree DNA.
Whilst other DNA companies do not offer a separate Y-DNA test examining STRs and matching, some do provide the Y-haplogroup a part of their single combined DNA test: Living DNA (around 2,000 SNPs) and 23andMe around 20,000 SMPs) (data source: ISOGG wiki, Y-DNA SNP testing chart).
Remember
Using DNA testing to solve genealogical problems is dependent on the databases of test results. If no one else from your paternal line has yet tested, you won’t get any matches. In addition, all of the commercial companies databases currently contain more results of those from the US than anywhere else. This is changing as more and more test, from all over the world. The test results are powerful but you may have to be patient.
A plea from me
Do you have any COWLING ancestors from England? I have registered a Cowling One Name Study and am interested in expanding this to incorporate a DNA Project. If you are a male Cowling descended from Cowlings in England, particularly those from Cambridgeshire, Yorkshire and Cornwall and would be interested in taking part do please get in touch. Similarly if you have Cowling relatives and are just interested in the One Name Study, do get in touch too. I would love to hear from you.
Up next
The next post will focus on mtDNA (mitochondrial DNA), the DNA we can use to assess ancient origins on the female line.