Demystifying DNA 3: mitochondrial DNA (mtDNA) tests
Originally posted 2nd August 2018, updated 4th October 2024.
This is the third in my series of posts attempting to demystify DNA testing for family historians. If you would like an overview to the many types of DNA test, do see the introduction to DNA testing post. In my last blog we looked at Y-DNA testing in more detail: when to use Y-DNA, who can test, where to test and how to interpret the results.
This month we move onto mitochondrial DNA, or mtDNA, testing.
Uses of mitochondrial DNA (mtDNA)
Mitochondrial DNA is to maternal research what Y-DNA testing is to paternal research as illustrated by the following schematic:
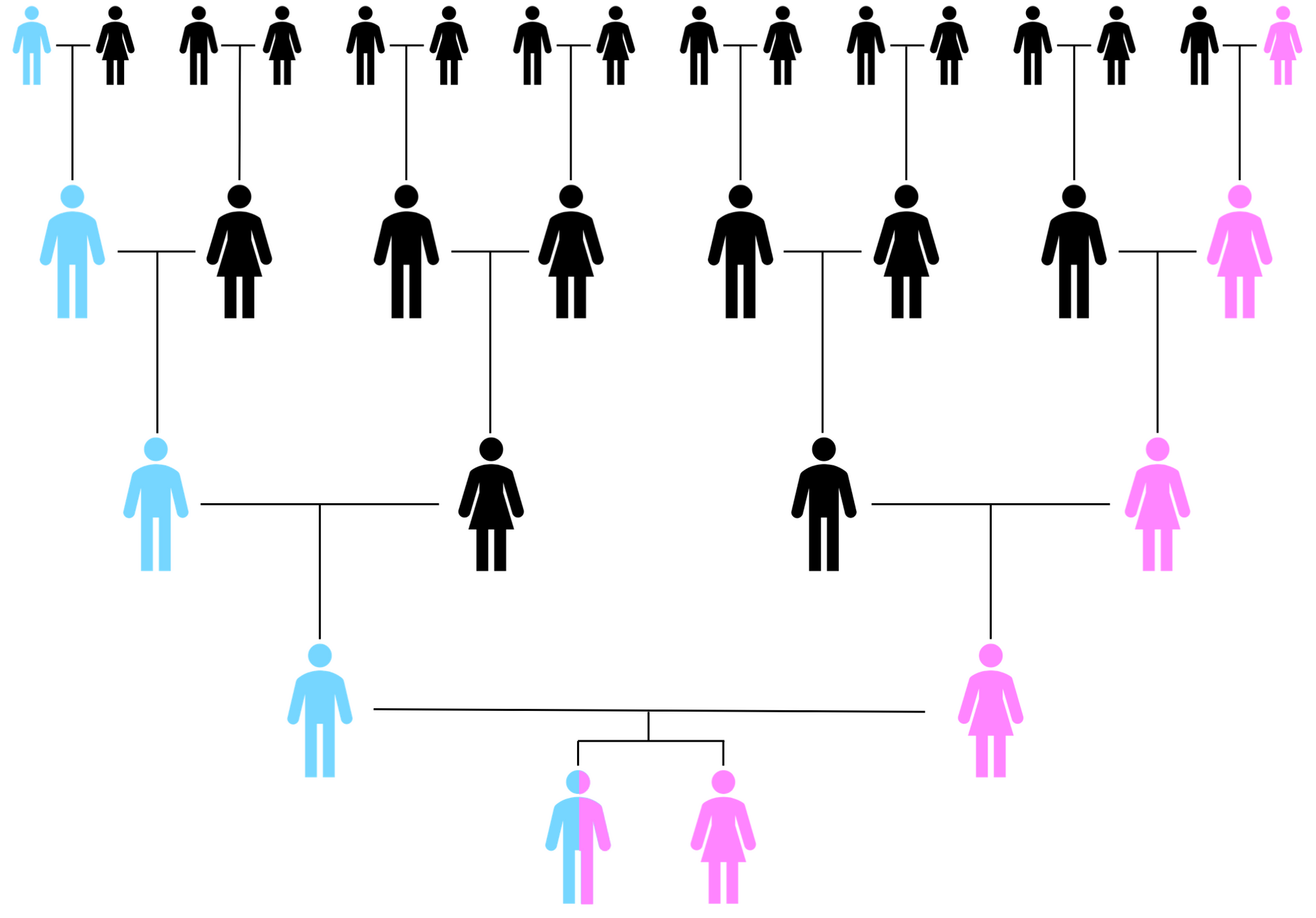
mtDNA is specific to the matrilineal (female to female) line, looking at your mother, her mother, her mother’s mother and so on. It does not include ALL of your mother’s ancestors, just those highlighted above. The difference to Y-DNA however, is that everyone has mtDNA. A mother passes on mtDNA to all of her children, males and females, but only females pass it on to the next generation. So a male can take a mtDNA test to find out about his matrilineal line.
Just like Y-DNA, mtDNA is passed largely unchanged from one generation to another, enabling its use for tracing maternal ancient origins. You can also, in theory, use mtDNA to support your genealogy research. However this is complicated by the fact that the surname changes at every generation, making traditional research more challenging, and the way in which mtDNA mutates.
Every now and again a mutation or copying error occurs with mtDNA, moving from one generation to the next. When I use the term “mutation” here I simply mean a change in DNA, no implication of anything to do with health. The main difference between the inheritance of mtDNA compared to Y-DNA is that the mutation rate of mtDNA is slower. A mtDNA match could share a common ancestor with you in recent generations or hundreds or even thousands of years ago.
mtDNA is therefore not as useful for testing speculatively to find matches, as it is less likely that a mtDNA match will share with you a common ancestor in a genealogically relevant timeframe. The beauty of a match on mtDNA is the fact that you will know what small section of your family tree the match is connected to. Where mtDNA is particularly useful is for confirming suspected relationships. It is a very powerful test for comparing your own data with a suspected match to see if you are indeed related to the same maternal ancestor. This approach can equally be applied to adoption cases as to the situation where you have two candidates for your maternal great grandmother.
Who can take a test?
Contrary to popular belief, mtDNA tests can actually be taken by both males and females. mtDNA passes from a mother to her children, the difference being that only the females then pass this mtDNA on. This is illustrated more clearly by the following schematic:
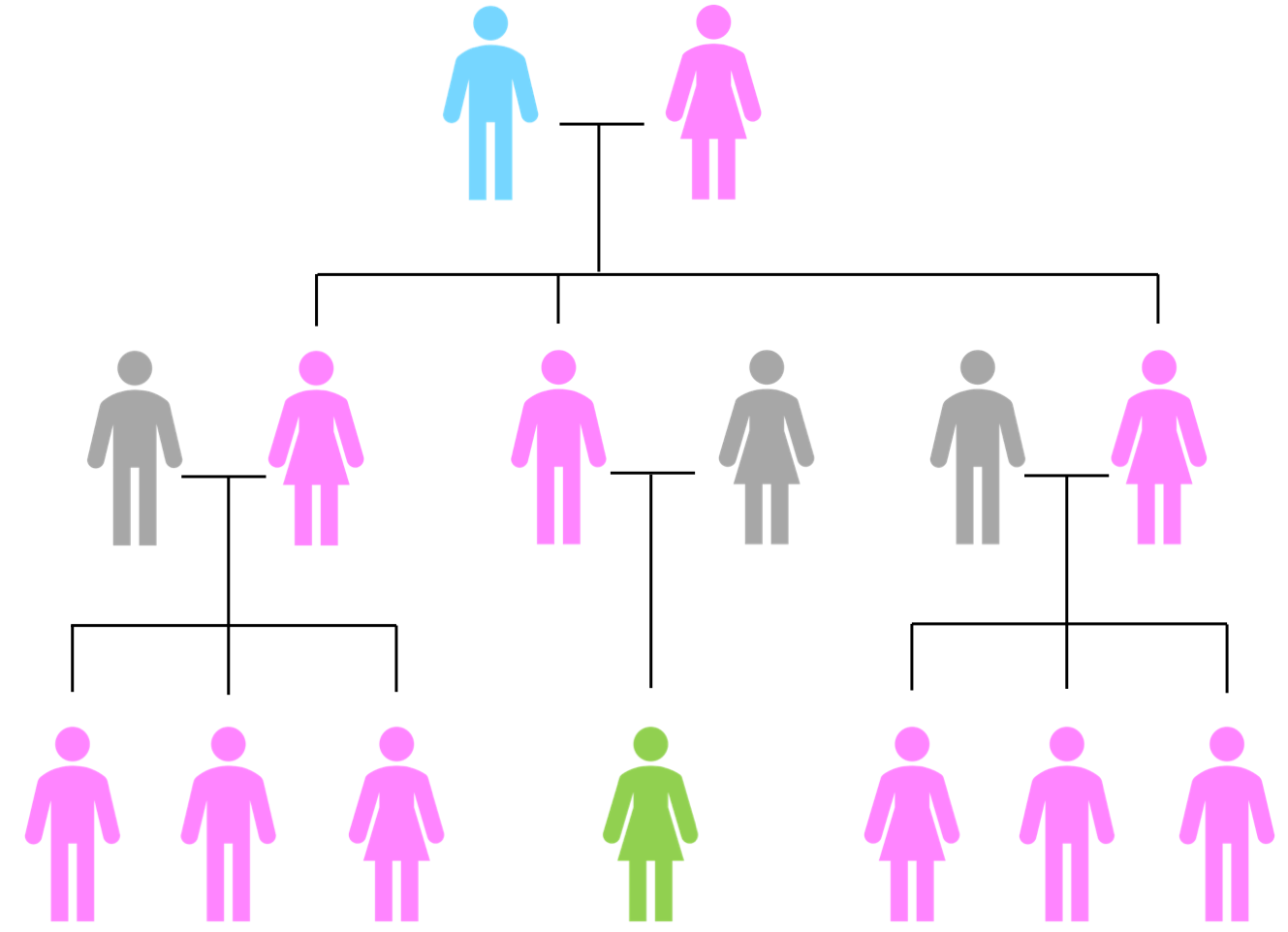
In the diagram, the descendants of a couple are shown. They have three children: two girls and a boy. The spouses of these three children are shown in grey to indicate they are not blood relatives of the couple at the top. All of the descendants of the uppermost female, who carry her mtDNA are shown in pink. This means that there are some females and some males coloured in pink. In the middle is a female filled in in green. Her father received mtDNA from his mother, but males do not pass on mtDNA to their children. The lady in green therefore does not carry her paternal grandmother's mtDNA.
Let's assume that this green female is the genealogist of the family and she wants to know more about her paternal grandmother's matrilineal (female to female) line. Unfortunately, her grandmother died some years ago. Looking at the diagram you can see that she has options to ask not only her aunts and first cousins, but also her father could take a mtDNA test to represent her grandmother's mtDNA. What if there are no surviving relatives to test? Just like with Y-DNA, mtDNA remains unchanged for many generations, so you could go back another generation to our genealogist's great grandmother, and look at other any female to female line descendants in the form of second cousins, and so on.
Generally speaking, if you have a choice, test the older generation. This approach does assume that all those in pink are the blood line of the uppermost female. A misattributed paternity event is a possibility and you should always test more than one candidate from more than one line of descent if you can.
Types of Test
Mitochondrial DNA is a circle of DNA, consisting of 16,569 base pairs (see the first post in this series, the introduction to DNA, for an explanation of the terminology). Mitochondrial DNA consists of the following regions:
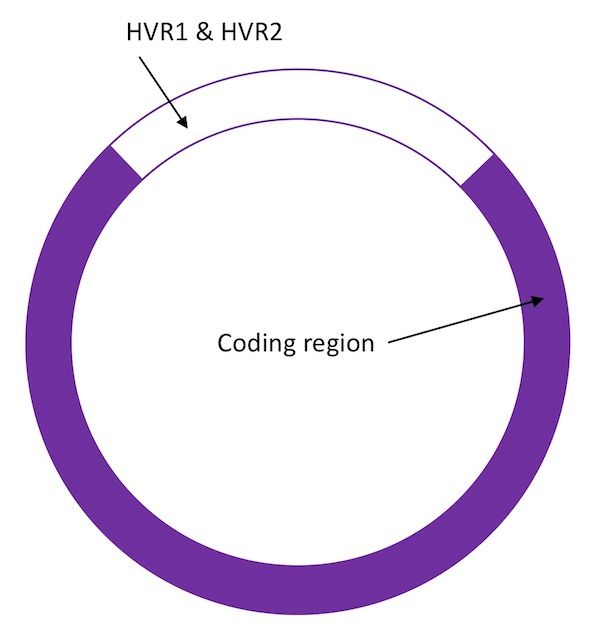
The area shown in white represents the hyper variable control regions (HVR1 & HVR2). These are the areas of the mtDNA known to mutate more quickly. They are therefore more likely to differ from one individual to another, unless they are closely related. The coding region undergoes changes less frequently.
The first mtDNA tests analysed DNA in the HVR1 and HVR2 control regions only. Later mtDNA tests included both the HVR1 & HVR2 regions and the coding region. Testing originally used SNP testing. Remember from the piece on Y-DNA testing, a Single Nucleotide Polymorph, or SNP, is a point along the DNA molecule known to differ from one individual to another – a point at which a mutation has occurred at some point in time. SNP (pronounced “snip”) testing analyses which nucleotide is found at many individual locations or SNPs.
Also available for mtDNA are sequence tests. Rather than look at individual SNPs all base pairs are analysed in the region of interest. Early tests just looked at the base pairs in the HVR1 or HVR1 and HVR2 regions. Now the standard is a full sequence test, which analyses all 16,569 base pairs. In much the same way as a higher number of markers on a Y-DNA STR tests gives you better data for comparison with others, more accurate mtDNA data is found with a full sequence test.
If we imagine the ring of DNA opened out flat then a visual representation of the difference is:
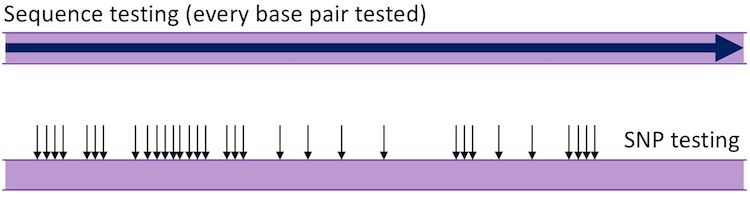
The Data
When we looked at the Y-DNA STR tests we looked directly at the number of repeats at STR markers or the identity of bases at particular locations. mtDNA data analysis is different. Here we compare how each individual differs from reference standards. The first produced was the Cambridge Reference Standard (CRS), now superseded by the corrected revised Cambridge Reference Standard (rCRS), based on a European who had haplogroup H. A second standard, the Reconstructed Sapiens Reference Sequence (RSRS), was produced more recently and was an attempt to compare mtDNA against a reference with an older haplogroup, closer to Mitochondrial Eve (see below for more on haplogroups). The details of the two standards are not appropriate here, more information can be found at the ISOGG website. It is, however, important to know which standard has been used by your testing company of choice if you are to compare results with those obtained elsewhere.
Family Tree DNA supplies results against both reference standards. The images below show the (truncated) results of my own mtDNA test against the rCRS at Family Tree DNA.
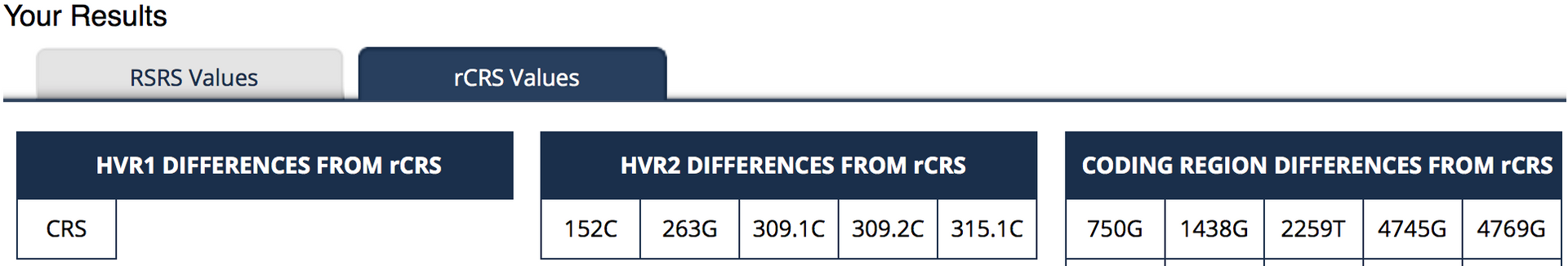

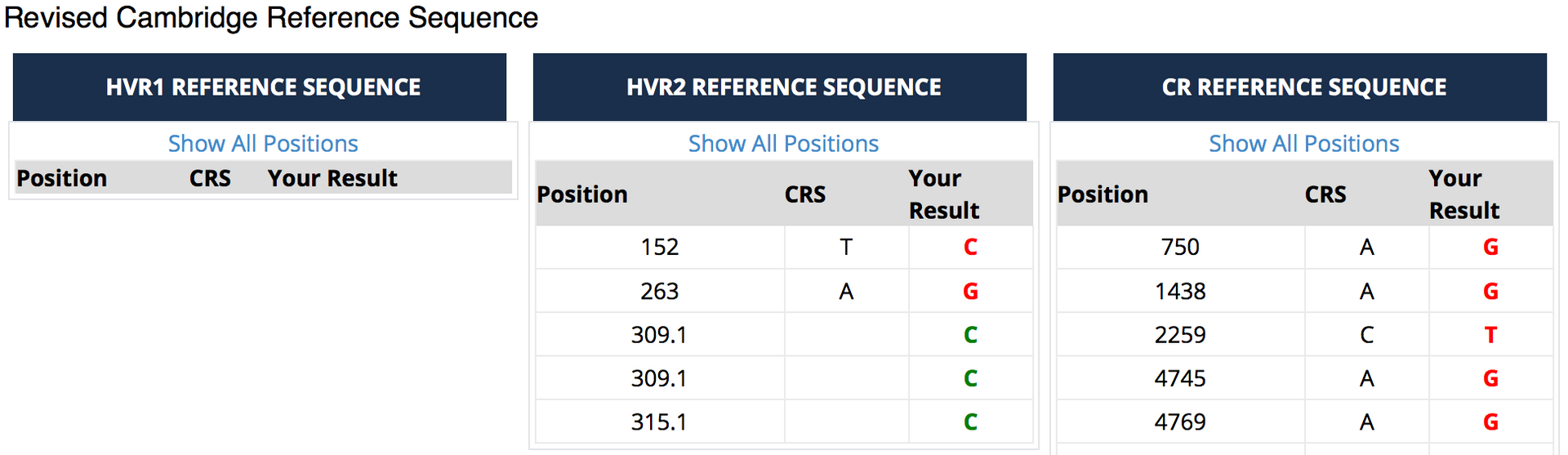
The results are actually reported in two ways, just to confuse you! In this case there are no differences to the standard in the HVR1 region. In the HVR2 region five differences are shown. The traditional way of reporting these is to the give the position number, followed by the letter of the base that you have compared to the original. So at position 152 I have C instead of the base of the rCRS. The second set of data (the lower table labelled “Revised Cambridge Reference Standard”) actually shows this more simply. It shows you that there should be a T at position 152 but I have a C.
The addition of a “.1” indicates an addition at this position. In fact I have two additional Cs at position 309. If a base is missing at a particular position it would be marked e.g. 309-, known as a deletion.
Now let’s turn our attention to the RSRS results, again my own (truncated) data:
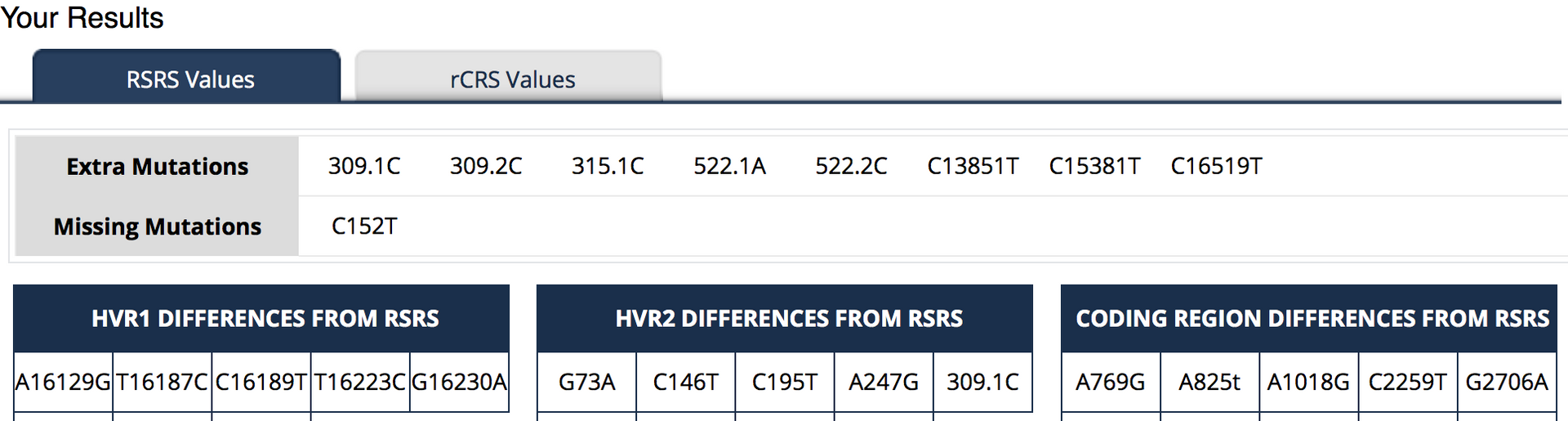
There are some differences against the reference standard in the HVR1 region here. This is to be expected: The reference for the rCRS was in haplogroup H, as am I, whereas the reference for RSRS is based on older haplogroups. Here differences are marked:
<reference base> POSITION NUMBER <your result>
so you can readily compare the base in the reference standard with your own. For the RSRS results there are also extra mutations and missing mutations. These refer to differences from what is expected for my haplogroup compared to the RSRS.
My current matches on mtDNA are shown below:
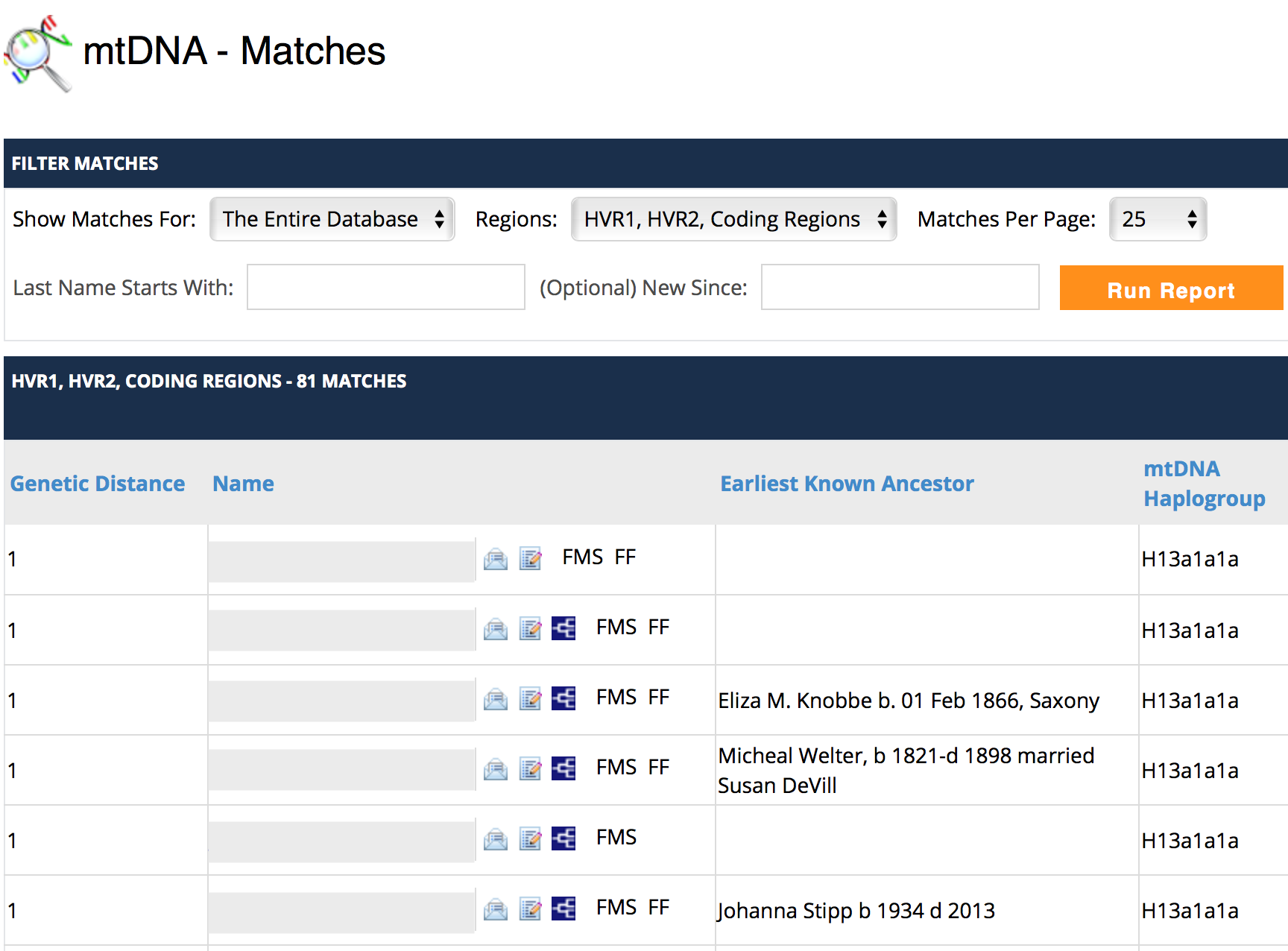
As you can see, I don’t yet have any matches at genetic distance of zero (this post was original published in 2018 and now, in 2024, I still do not have any zero genetic distance matches!). A genetic difference of 1 means that there is a difference in my data compared to the other test taker’s data at one position, whether it be a different base, an addition or a missing mutation compared to their results.
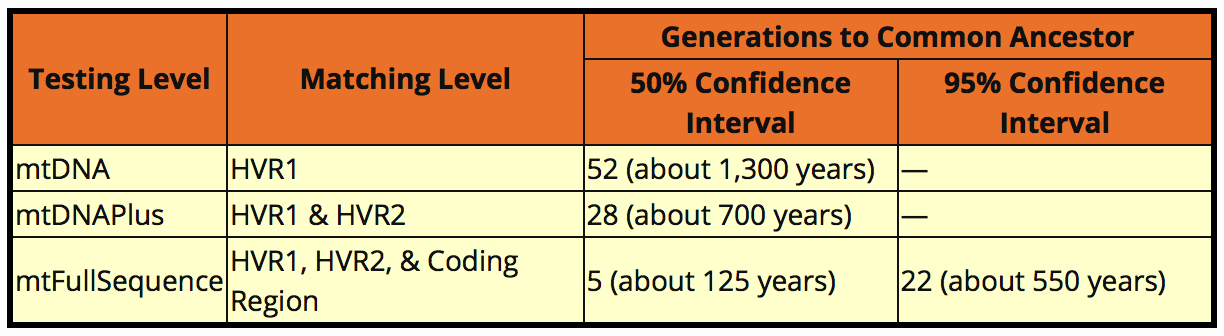
With Y-DNA we could calculate a reasonable estimate of the time to Most Recent Common Ancestor (MRCA) as Y-DNA mutations happen at a regular rate and there is some level of confidence in predictability. As I said earlier, with mtDNA the mutation rates are much slower and there is much greater range. The following table is taken from the Family Tree DNA website. Even if I had a match with a genetic distance of zero there’s only a 50% likelihood that person and I share an ancestor within 5 generations. It’s more likely that the common ancestor is somewhere within the last 5-22 generations.
Mitochondrial DNA (mtDNA) haplogroups
What I find interesting is knowledge of my mtDNA haplogroup. Just as there is a haplogroup tree for Y-DNA, there is an equivalent mtDNA haplogroup tree, as all females are descended from mitochondrial Eve. An individual’s mtDNA haplogroup is their location in the human mtDNA haplogroup tree. Everyone fits on this tree, some branches dating far further back in time than those derived from more recent mutations. A simple graphic is shown below but there are many branches, or subclades, within each haplogroup.
Each haplogroup is connected to particular time and place and more information on where the haplogroups originated can be found here: mtDNA haplogroups. My own haplogroup is H. This is a predominantly European haplogroup as I would expect and does not reveal anything exciting about my own family history. However, for those with a family story that 3x great grandmother was a local Indian girl that 3 x great grandfather met while he worked in British India, discovering the haplogroup can be very important.
The Testing Companies
I’m only considering the main five DNA testing companies in this series of blogs, to keep things simple. Of these, only Family Tree DNA currently offers separate mtDNA tests with matching and, in 2024 only the full sequence test remains available.
Whilst the other DNA companies in “the big five” do not offer a separate mtDNA tests, both Living DNA and 23 and Me includes measurement of 4000+ positions on the mtDNA genome to define the haplogroup (data source: ISOGG wiki, MtDNA testing comparison chart).
Remember
Be careful with haplogroups – Y-DNA and mtDNA lettering conventions do not relate to one another. The Y-DNA haplogroup is an indication of paternal origins, the mtDNA haplogroup an indication of maternal origins. A man has both, a woman has only a mtDNA haplogroup.
As with all DNA tests, the number of matches you get with mtDNA testing will depend on who else has tested. If you have no matches to start with: be patient.
With any type of DNA test, the results obtained form only part of the analysis. DNA testing does not answer questions alone: it must always be assessed along with other information and documentary evidence.
Up Next
The next post will focus on the most popular type of DNA testing now: autosomal DNA, the type of test offered by Ancestry, My Heritage and Family Tree DNA (the Family Finder test) to find matches to close living relatives.